aonori3/symbia_covid_20figs 🖼️🔢❓📝✓ → 🖼️
About
Example Output
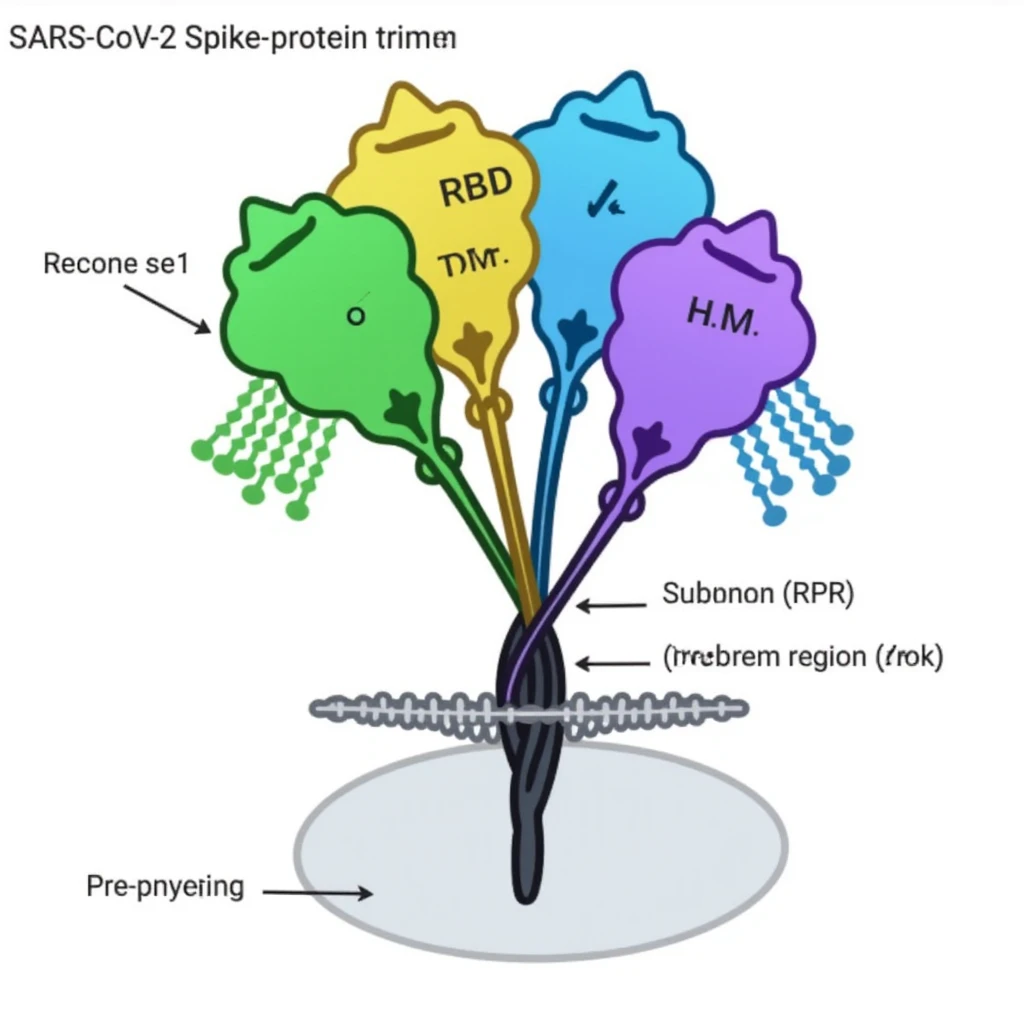
"covidfig of SARS-CoV-2 spike protein trimer in its prefusion state. 3 distinct protomers forming a mushroom-like structure, each protomer highlighted in a different color: one in light blue, one in light green, one in light purple. The spike protein consists of an S1 subunit (larger top portion) and an S2 subunit (lower stem portion). S1 subunits include receptor-binding domains (RBDs), one in 'up' conformation colored in bright yellow to indicate an active binding state, two in 'down' conformation in light blue and purple respectively. Glycans depicted as green tree-like structures distributed along the surface of each protomer. S2 subunit forming the elongated stalk is represented with a textured cylindrical structure to emphasize heptad repeats and fusion peptide regions. Membrane-proximal region (MPR) shown in dark grey near the base, indicating proximity to the viral envelope. Overall, structure sits atop a semi-transparent grey circle representing the viral membrane. Labels for RBD, NTD (N-terminal domain in the S1 subunit), HR1/HR2 (heptad repeat regions), and fusion peptide included for clarity."
Output

Performance Metrics
All Input Parameters
{
"model": "dev",
"prompt": "covidfig of SARS-CoV-2 spike protein trimer in its prefusion state. 3 distinct protomers forming a mushroom-like structure, each protomer highlighted in a different color: one in light blue, one in light green, one in light purple. The spike protein consists of an S1 subunit (larger top portion) and an S2 subunit (lower stem portion). S1 subunits include receptor-binding domains (RBDs), one in 'up' conformation colored in bright yellow to indicate an active binding state, two in 'down' conformation in light blue and purple respectively. Glycans depicted as green tree-like structures distributed along the surface of each protomer. S2 subunit forming the elongated stalk is represented with a textured cylindrical structure to emphasize heptad repeats and fusion peptide regions. Membrane-proximal region (MPR) shown in dark grey near the base, indicating proximity to the viral envelope. Overall, structure sits atop a semi-transparent grey circle representing the viral membrane. Labels for RBD, NTD (N-terminal domain in the S1 subunit), HR1/HR2 (heptad repeat regions), and fusion peptide included for clarity.",
"lora_scale": 1,
"num_outputs": 1,
"aspect_ratio": "1:1",
"output_format": "webp",
"guidance_scale": 3.5,
"output_quality": 90,
"prompt_strength": 0.8,
"extra_lora_scale": 1,
"num_inference_steps": 28
}
Input Parameters
- mask
- Image mask for image inpainting mode. If provided, aspect_ratio, width, and height inputs are ignored.
- seed
- Random seed. Set for reproducible generation
- image
- Input image for image to image or inpainting mode. If provided, aspect_ratio, width, and height inputs are ignored.
- model
- Which model to run inference with. The dev model performs best with around 28 inference steps but the schnell model only needs 4 steps.
- width
- Width of generated image. Only works if `aspect_ratio` is set to custom. Will be rounded to nearest multiple of 16. Incompatible with fast generation
- height
- Height of generated image. Only works if `aspect_ratio` is set to custom. Will be rounded to nearest multiple of 16. Incompatible with fast generation
- prompt (required)
- Prompt for generated image. If you include the `trigger_word` used in the training process you are more likely to activate the trained object, style, or concept in the resulting image.
- go_fast
- Run faster predictions with model optimized for speed (currently fp8 quantized); disable to run in original bf16
- extra_lora
- Load LoRA weights. Supports Replicate models in the format <owner>/<username> or <owner>/<username>/<version>, HuggingFace URLs in the format huggingface.co/<owner>/<model-name>, CivitAI URLs in the format civitai.com/models/<id>[/<model-name>], or arbitrary .safetensors URLs from the Internet. For example, 'fofr/flux-pixar-cars'
- lora_scale
- Determines how strongly the main LoRA should be applied. Sane results between 0 and 1 for base inference. For go_fast we apply a 1.5x multiplier to this value; we've generally seen good performance when scaling the base value by that amount. You may still need to experiment to find the best value for your particular lora.
- megapixels
- Approximate number of megapixels for generated image
- num_outputs
- Number of outputs to generate
- aspect_ratio
- Aspect ratio for the generated image. If custom is selected, uses height and width below & will run in bf16 mode
- output_format
- Format of the output images
- guidance_scale
- Guidance scale for the diffusion process. Lower values can give more realistic images. Good values to try are 2, 2.5, 3 and 3.5
- output_quality
- Quality when saving the output images, from 0 to 100. 100 is best quality, 0 is lowest quality. Not relevant for .png outputs
- prompt_strength
- Prompt strength when using img2img. 1.0 corresponds to full destruction of information in image
- extra_lora_scale
- Determines how strongly the extra LoRA should be applied. Sane results between 0 and 1 for base inference. For go_fast we apply a 1.5x multiplier to this value; we've generally seen good performance when scaling the base value by that amount. You may still need to experiment to find the best value for your particular lora.
- replicate_weights
- Load LoRA weights. Supports Replicate models in the format <owner>/<username> or <owner>/<username>/<version>, HuggingFace URLs in the format huggingface.co/<owner>/<model-name>, CivitAI URLs in the format civitai.com/models/<id>[/<model-name>], or arbitrary .safetensors URLs from the Internet. For example, 'fofr/flux-pixar-cars'
- num_inference_steps
- Number of denoising steps. More steps can give more detailed images, but take longer.
- disable_safety_checker
- Disable safety checker for generated images.
Output Schema
Output
Example Execution Logs
Using seed: 43569 Prompt: covidfig of SARS-CoV-2 spike protein trimer in its prefusion state. 3 distinct protomers forming a mushroom-like structure, each protomer highlighted in a different color: one in light blue, one in light green, one in light purple. The spike protein consists of an S1 subunit (larger top portion) and an S2 subunit (lower stem portion). S1 subunits include receptor-binding domains (RBDs), one in 'up' conformation colored in bright yellow to indicate an active binding state, two in 'down' conformation in light blue and purple respectively. Glycans depicted as green tree-like structures distributed along the surface of each protomer. S2 subunit forming the elongated stalk is represented with a textured cylindrical structure to emphasize heptad repeats and fusion peptide regions. Membrane-proximal region (MPR) shown in dark grey near the base, indicating proximity to the viral envelope. Overall, structure sits atop a semi-transparent grey circle representing the viral membrane. Labels for RBD, NTD (N-terminal domain in the S1 subunit), HR1/HR2 (heptad repeat regions), and fusion peptide included for clarity. [!] txt2img mode Using dev model free=8756963061760 Downloading weights 2024-10-03T20:41:06Z | INFO | [ Initiating ] chunk_size=150M dest=/tmp/tmp21hvpbtr/weights url=https://replicate.delivery/yhqm/zAf2PS8nXhX0WqlHAWvlWkP6apflTYDLcjffPIsvrBc3tsMOB/trained_model.tar 2024-10-03T20:41:07Z | INFO | [ Complete ] dest=/tmp/tmp21hvpbtr/weights size="172 MB" total_elapsed=1.740s url=https://replicate.delivery/yhqm/zAf2PS8nXhX0WqlHAWvlWkP6apflTYDLcjffPIsvrBc3tsMOB/trained_model.tar Downloaded weights in 1.77s Loaded LoRAs in 2.50s 0%| | 0/28 [00:00<?, ?it/s] 4%|▎ | 1/28 [00:00<00:09, 2.88it/s] 7%|▋ | 2/28 [00:00<00:08, 3.21it/s] 11%|█ | 3/28 [00:00<00:08, 3.05it/s] 14%|█▍ | 4/28 [00:01<00:08, 2.98it/s] 18%|█▊ | 5/28 [00:01<00:07, 2.94it/s] 21%|██▏ | 6/28 [00:02<00:07, 2.92it/s] 25%|██▌ | 7/28 [00:02<00:07, 2.91it/s] 29%|██▊ | 8/28 [00:02<00:06, 2.90it/s] 32%|███▏ | 9/28 [00:03<00:06, 2.89it/s] 36%|███▌ | 10/28 [00:03<00:06, 2.89it/s] 39%|███▉ | 11/28 [00:03<00:05, 2.88it/s] 43%|████▎ | 12/28 [00:04<00:05, 2.88it/s] 46%|████▋ | 13/28 [00:04<00:05, 2.88it/s] 50%|█████ | 14/28 [00:04<00:04, 2.88it/s] 54%|█████▎ | 15/28 [00:05<00:04, 2.88it/s] 57%|█████▋ | 16/28 [00:05<00:04, 2.88it/s] 61%|██████ | 17/28 [00:05<00:03, 2.88it/s] 64%|██████▍ | 18/28 [00:06<00:03, 2.88it/s] 68%|██████▊ | 19/28 [00:06<00:03, 2.88it/s] 71%|███████▏ | 20/28 [00:06<00:02, 2.88it/s] 75%|███████▌ | 21/28 [00:07<00:02, 2.88it/s] 79%|███████▊ | 22/28 [00:07<00:02, 2.88it/s] 82%|████████▏ | 23/28 [00:07<00:01, 2.88it/s] 86%|████████▌ | 24/28 [00:08<00:01, 2.88it/s] 89%|████████▉ | 25/28 [00:08<00:01, 2.88it/s] 93%|█████████▎| 26/28 [00:08<00:00, 2.88it/s] 96%|█████████▋| 27/28 [00:09<00:00, 2.88it/s] 100%|██████████| 28/28 [00:09<00:00, 2.88it/s] 100%|██████████| 28/28 [00:09<00:00, 2.90it/s]
Version Details
- Version ID
b1d207de93444dcb7d9e65e616ec7906c865b9afdf3b230b61fba2c15198817f- Version Created
- October 3, 2024